- Scientific Journals (peer reviewed articles)
- Book Chapters
- International Conferences (peer reviewed articles)
- International Conferences (invited oral presentation)
- International Conferences (abstracts)
- International Conferences (other presentations)
- National Conferences
- Postgraduate Dissertations
Scientific Journals (peer reviewed articles)
| [10] |
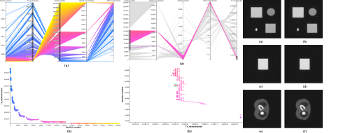 Cameron C. Gray, Shatha F. Al-Maliki, & Franck P. Vidal.
Data exploration in evolutionary reconstruction of PET images.
Genetic Programming and Evolvable Machines, 19(3):391-419,
September 2018.
Cameron C. Gray, Shatha F. Al-Maliki, & Franck P. Vidal.
Data exploration in evolutionary reconstruction of PET images.
Genetic Programming and Evolvable Machines, 19(3):391-419,
September 2018.



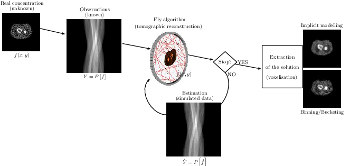
This work is based on a cooperative co-evolution algorithm called `Fly Algorithm', which is an evolutionary algorithm (EA) where individuals are called `flies'. It is a specific case of the `Parisian Approach' where the solution of an optimisation problem is a set of individuals (e.g. the whole population) instead of a single individual (the best one) as in typical EAs. The optimisation problem considered here is tomography reconstruction in positron emission tomography (PET). It estimates the concentration of a radioactive substance (called a radiotracer) within the body. Tomography, in this context, is considered as a difficult ill-posed inverse problem. The Fly Algorithm aims at optimising the position of 3-D points that mimic the radiotracer. At the end of the optimisation process, the fly population is extracted as it corresponds to an estimate of the radioactive concentration. During the optimisation loop a lot of data is generated by the algorithm, such as image metrics, duration, and internal states. This data is recorded in a log file that can be post-processed and visualised. We propose using information visualisation and user interaction techniques to explore the algorithm's internal data. Our aim is to better understand what happens during the evolutionary loop. Using an example, we demonstrate that it is possible to interactively discover when an early termination could be triggered. It is implemented in a new stopping criterion. It is tested on two other examples on which it leads to a 60% reduction of the number of iterations without any loss of accuracy. Keywords: Fly Algorithm; Tomography reconstruction; Information visualisation; Data exploration; Artificial evolution; Parisian evolution. |
| [9] |
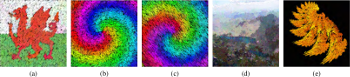 Z. Ali Abbood, and F. P. Vidal.
Fly4Arts: Evolutionary digital art with the Fly algorithm.
ISTE Arts & Science, 17(1):11--16, October 2017,
October 2017.
Z. Ali Abbood, and F. P. Vidal.
Fly4Arts: Evolutionary digital art with the Fly algorithm.
ISTE Arts & Science, 17(1):11--16, October 2017,
October 2017.



The aim of this study is to generate artistic images, such as digital mosaics, as an optimisation problem without the introduction of any a priori knowledge or constraint other than an input image. The usual practice to produce digital mosaic images heavily relies on Centroidal Voronoi diagrams. We demonstrate here that it can be modelled as an optimisation problem solved using a cooperative co-evolution strategy based on the Parisian evolution approach, the Fly algorithm. An individual is called a fly. Its aim of the algorithm is to optimise the position of infinitely small 3-D points (the flies). The Fly algorithm has been initially used in real-time stereo vision for robotics. It has also demonstrated promising results in image reconstruction for tomography. In this new application, a much more complex representation has been study. A fly is a tile. It has its own position, size, colour, and rotation angle. Our method takes advantage of graphics processing units (GPUs) to generate the images using the modern OpenGL Shading Language (GLSL) and Open Computing Language (OpenCL) to compute the difference between the input image and simulated image. Different types of tiles are implemented, some with transparency, to generate different visual effects, such as digital mosaic and spray paint. An online study with 41 participants has been conducted to compare some of our results with those generated using an open-source software for image manipulation. It demonstrates that our method leads to more visually appealing images. Keywords: Digital mosaic; Evolutionary art; Fly Algorithm; Parisian evolution; Cooperative co-evolution |
| [8] |
 Z. Ali Abbood, J. Lavauzelle, É Lutton, J.-M. Rocchisani, J. Louchet, and F. P. Vidal.
Voxelisation in the 3-D Fly Algorithm for PET.
Swarm and Evolutionary Computation, 36:91-105,
October 2017.
Z. Ali Abbood, J. Lavauzelle, É Lutton, J.-M. Rocchisani, J. Louchet, and F. P. Vidal.
Voxelisation in the 3-D Fly Algorithm for PET.
Swarm and Evolutionary Computation, 36:91-105,
October 2017.



The Fly Algorithm was initially developed for 3-D robot vision applications. It consists in solving the inverse problem of shape reconstruction from projections by evolving a population of 3-D points in space (the `flies'), using an evolutionary optimisation strategy. Here, in its version dedicated to tomographic reconstruction in medical imaging, the flies are mimicking radioactive photon sources. Evolution is controlled using a fitness function based on the discrepancy of the projections simulated by the flies with the actual pattern received by the sensors. The reconstructed radioactive concentration is derived from the population of flies, i.e. a collection of points in the 3-D Euclidean space, after convergence. `Good' flies were previously binned into voxels. In this paper, we study which flies to include in the final solution and how this information can be sampled to provide more accurate datasets in a reduced computation time. We investigate the use of density fields, based on Metaballs and on Gaussian functions respectively, to obtain a realistic output. The spread of each Gaussian kernel is modulated in function of the corresponding fly fitness. The resulting volumes are compared with previous work in terms of normalised-cross correlation. In our test-cases, data fidelity increases by more than 10% when density fields are used instead of binning. Our method also provides reconstructions comparable to those obtained using well-established techniques used in medicine (filtered back- projection and ordered subset expectation-maximisation). Keywords: Fly algorithm; Evolutionary computation; tomography reconstruction; iterative algorithms; inverse problems; co-operative co-evolution |
| [7] |
F. P. Vidal, and P. F. Villard.
Development and validation of real-time simulation of X-ray imaging
with respiratory motion.
Computerized Medical Imaging and Graphics, 49:1-15, April 2016.




We present a framework that combines evolutionary optimisation, soft tissue modelling and ray tracing on GPU to simultaneously compute the respiratory motion and X-ray imaging in real-time. Our aim is to provide validated building blocks with high fidelity to closely match both the human physiology and the physics of X-rays. A CPU-based set of algorithms is presented to model organ behaviours during respiration. Soft tissue deformation is computed with an extension of the Chain Mail method. Rigid elements move according to kinematic laws. A GPU-based surface rendering method is proposed to compute the X-ray image using the Beer–Lambert law. It is provided as an open-source library. A quantitative validation study is provided to objectively assess the accuracy of both components: (i) the respiration against anatomical data, and (ii) the X-ray against the Beer–Lambert law and the results of Monte Carlo simulations. Our implementation can be used in various applications, such as interactive medical virtual environment to train percutaneous transhepatic cholangiography in interventional radiology, 2D/3D registration, computation of digitally reconstructed radiograph, simulation of 4D sinograms to test tomography reconstruction tools. Keywords: X-ray simulation, Deterministic simulation (ray-tracing), Digitally reconstructed radiograph, Respiration simulation, Medical virtual environment, Imaging guidance, Interventional radiology training |
| [6] |
 P. F. Villard, F. P. Vidal, L. ap Cenydd, R. Holbrey, S. Pisharody,
S. Johnson, A. Bulpitt, N. W. John, F. Bello, and D. Gould.
Interventional radiology virtual simulator for liver biopsy.
International Journal of Computer Assisted Radiology and
Surgery, 9(2):255-267, March 2014.
P. F. Villard, F. P. Vidal, L. ap Cenydd, R. Holbrey, S. Pisharody,
S. Johnson, A. Bulpitt, N. W. John, F. Bello, and D. Gould.
Interventional radiology virtual simulator for liver biopsy.
International Journal of Computer Assisted Radiology and
Surgery, 9(2):255-267, March 2014.




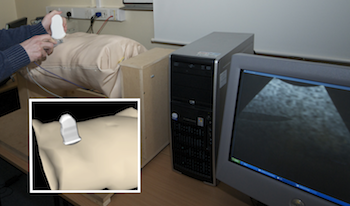
Purpose: Training in Interventional Radiology currently uses the apprenticeship model, where clinical and technical skills of invasive procedures are learnt during practice in patients. This apprenticeship training method is increasingly limited by regulatory restrictions on working hours, concerns over patient risk through trainees’ inexperience and the variable exposure to case mix and emergencies during training. To address this, we have developed a computer-based simulation of visceral needle puncture procedures. Methods: A real-time framework has been built that includes: segmentation, physically based modelling, haptics rendering, pseudo-ultrasound generation and the concept of a physical mannequin. It is the result of a close collaboration between different universities, involving computer scientists, clinicians, clinical engineers and occupational psychologists. Results: The technical implementation of the framework is a robust and real-time simulation environment combining a physical platform and an immersive computerized virtual environment. The face, content and construct validation have been previously assessed, showing the reliability and effectiveness of this framework, as well as its potential for teaching visceral needle puncture. Conclusion: A simulator for ultrasound-guided liver biopsy has been developed. It includes functionalities and metrics extracted from cognitive task analysis. This framework can be useful during training, particularly given the known difficulties in gaining significant practice of core skills in patients. Keywords: Biomedical computing, Image segmentation, Simulation, Virtual reality |
| [5] |
 F. P. Vidal, P.-F. Villard, and É. Lutton.
Tuning of patient specific deformable models using an adaptive
evolutionary optimization strategy.
IEEE Transactions on Biomedical Engineering, 59(10):2942-2949,
October 2012.
F. P. Vidal, P.-F. Villard, and É. Lutton.
Tuning of patient specific deformable models using an adaptive
evolutionary optimization strategy.
IEEE Transactions on Biomedical Engineering, 59(10):2942-2949,
October 2012.




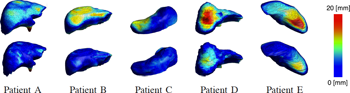
We present and analyze the behavior of an evolutionary algorithm designed to estimate the parameters of a complex organ behavior model. The model is adaptable to account for patients specificities. The aim is to finely tune the model to be accurately adapted to various real patient datasets. It can then be embedded, for example, in high fidelity simulations of the human physiology. We present here an application focused on respiration modeling. The algorithm is automatic and adaptive. A compound fitness function has been designed to take into account for various quantities that have to be minimized. The algorithm efficiency is experimentally analyzed on several real test-cases: i) three patient datasets have been acquired with the breath hold protocol, and ii) two datasets corresponds to 4D CT scans. Its performance is compared with two traditional methods (downhill simplex and conjugate gradient descent), a random search and a basic realvalued genetic algorithm. The results show that our evolutionary scheme provides more significantly stable and accurate results. Keywords: Evolutionary computation, inverse problems, medical simulation, adaptive algorithm |
| [4] |
 P.-F. Villard, F. P. Vidal, C. Hunt, F. Bello, N. W. John, S. Johnson, and
D. A. Gould.
A prototype percutaneous transhepatic cholangiography training simulator with real-time breathing motion
International Journal of Computer Assisted Radiology and
Surgery, 4(6):571-578, November 2009.
P.-F. Villard, F. P. Vidal, C. Hunt, F. Bello, N. W. John, S. Johnson, and
D. A. Gould.
A prototype percutaneous transhepatic cholangiography training simulator with real-time breathing motion
International Journal of Computer Assisted Radiology and
Surgery, 4(6):571-578, November 2009.




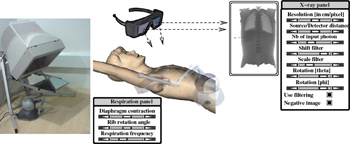
Purpose: We present here a simulator for interventional radiology focusing on percutaneous transhepatic cholangiography (PTC). This procedure consists of inserting a needle into the biliary tree using fluoroscopy for guidance. Methods: The requirements of the simulator have been driven by a task analysis. The three main components have been identified: the respiration, the real-time X-ray display (fluoroscopy) and the haptic rendering (sense of touch). The framework for modelling the respiratory motion is based on kinematics laws and on the Chainmail algorithm. The fluoroscopic simulation is performed on the graphic card and makes use of the Beer-Lambert law to compute the X-ray attenuation. Finally, the haptic rendering is integrated to the virtual environment and takes into account the soft-tissue reaction force feedback and maintenance of the initial direction of the needle during the insertion. Results: Five training scenarios have been created using patient-specific data. Each of these provides the user with variable breathing behaviour, fluoroscopic display tuneable to any device parameters and needle force feedback. Conclusions A detailed task analysis has been used to design and build the PTC simulator described in this paper. The simulator includes real-time respiratory motion with two independent parameters (rib kinematics and diaphragm action), on-line fluoroscopy implemented on the Graphics Processing Unit and haptic feedback to feel the soft-tissue behaviour of the organs during the needle insertion. Keywords: Interventional radiology; Virtual environments; Respiration simulation; X-ray simulation; Needle puncture; Haptics; Task analysis |
| [3] |
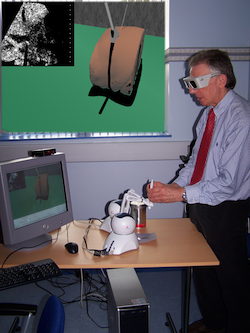 F. P. Vidal, N. W. John, D. A. Gould, and A. E. Healey.
Simulation of ultrasound guided needle puncture using patient
specific data with 3D textures and volume haptics.
Computer Animation and Virtual Worlds, 19(2):111-127, May
2008.
F. P. Vidal, N. W. John, D. A. Gould, and A. E. Healey.
Simulation of ultrasound guided needle puncture using patient
specific data with 3D textures and volume haptics.
Computer Animation and Virtual Worlds, 19(2):111-127, May
2008.



We present an integrated system for training ultrasound (US) guided needle puncture. Our aim is to provide a validated training tool for interventional radiology (IR) that uses actual patient data. IR procedures are highly reliant on the sense of touch and so haptic hardware is an important part of our solution. A hybrid surface/volume haptic rendering of an US transducer is proposed to constrain the device to remain outside the bony structures when scanning the patient's skin. A volume haptic model is proposed that implements an effective model of needle puncture. Force measurements have been made on real tissue and the resulting data is incorporated into the model. The other input data required is a computed tomography (CT) scan of the patient that is used to create the patient specific models. It is also the data source for a novel simulation of a virtual US scanner, which is used to guide the needle to the correct location. Keywords: medical virtual environment; imaging guidance; interventional radiology training; needle puncture; volume haptics; vertex and pixel shaders |
| [2] |
 F. P. Vidal, F. Bello, K. W. Brodlie, D. A. Gould, N. W. John, R. Phillips, and
N. J. Avis.
Principles and applications of computer graphics in medicine.
Computer Graphics Forum, 25(1):113-137, March 2006.
F. P. Vidal, F. Bello, K. W. Brodlie, D. A. Gould, N. W. John, R. Phillips, and
N. J. Avis.
Principles and applications of computer graphics in medicine.
Computer Graphics Forum, 25(1):113-137, March 2006.



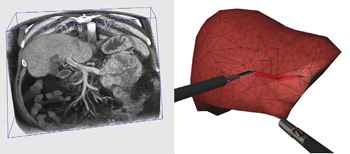
The medical domain provides excellent opportunities for the application of computer graphics, visualization and virtual environments, with the potential to help improve healthcare and bring benefits to patients. This survey paper provides a comprehensive overview of the state-of-the-art in this exciting field. It has been written from the perspective of both computer scientists and practising clinicians and documents past and current successes together with the challenges that lie ahead. The article begins with a description of the software algorithms and techniques that allow visualization of and interaction with medical data. Example applications from research projects and commercially available products are listed, including educational tools; diagnostic aids; virtual endoscopy; planning aids; guidance aids; skills training; computer augmented reality and use of high performance computing. The final section of the paper summarizes the current issues and looks ahead to future developments. Keywords: visualization; augmented and virtual realities; computer graphics; health; physically-based modeling; medical sciences; simulation |
| [1] |
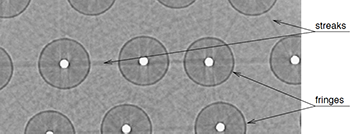 F. P. Vidal, J. M. Létang, G. Peix, and P. Clœtens.
Investigation of artefact sources in synchrotron microtomography via
virtual x-ray imaging.
Nuclear Instruments and Methods in Physics Research B,
234(3):333-348, June 2005.
F. P. Vidal, J. M. Létang, G. Peix, and P. Clœtens.
Investigation of artefact sources in synchrotron microtomography via
virtual x-ray imaging.
Nuclear Instruments and Methods in Physics Research B,
234(3):333-348, June 2005.



Qualitative and quantitative use of volumes reconstructed by computed tomography (CT) can be compromised due to artefacts which corrupt the data. This article illustrates a method based on virtual X-ray imaging to investigate sources of artefacts which occur in microtomography using synchrotron radiation. In this phenomenological study, different computer simulation methods based on physical X-ray properties, eventually coupled with experimental data, are used in order to compare artefacts obtained theoretically to those present in a volume acquired experimentally, or to predict them for a particular experimental setup. The article begins with the presentation of a synchrotron microtomographic slice of a reinforced fibre composite acquired at the European Synchrotron Radiation Facility (ESRF) containing streak artefacts. This experimental context is used as the motive throughout the paper to illustrate the investigation of some artefact sources. First, the contribution of direct radiation is compared to the contribution of secondary radiations. Then, the effect of some methodological aspects are detailed, including under-sampling, sample and camera misalignment, sample extending outside of the field of view and photonic noise. The effect of harmonic components present in the experimental spectrum are also simulated. Afterwards, detector properties, such as its impulse response or defective pixels, are taken into account. Finally, the importance of phase contrast effects is evaluated. In the last section, this investigation is discussed by putting emphasis on the experimental context which is used throughout this paper. Keywords: X-ray microtomography; Artefact; Deterministic simulation (ray-tracing); Monte Carlo method; Phase contrast; Modulation transfer function |
Book Chapters
| [2] |
F. P. Vidal, and J.-M. Rocchisani.
Graphics Processing Unit-Based High Performance Computing in Radiation Therapy, chapter Reconstruction in Positron Emission Tomography, pages 185-208.
CRC Press, 2015.
ISBN 9781482244793.


|
| [1] |
D. Gould, F. Bello, N. John, S. Johnson, C. Hunt, H. Woolnough, A. Bulpitt,
V. Luboz, D. King, P.-F. Villard, S. Pisharody, F. Vidal, and A. Sinha.
Innovative Cardiovascular Procedures, chapter Simulator
development in vascular and visceral interventions, pages 11-28.
Edizione Minerva Medica, Turin, Italy, 2009.
ISBN 88-7711-637-7.


Throughout the practice of procedural medicine, there is an unrelenting shift to management by less invasive techniques such as interventional radiology (IR). This subspecialty within radiology uses imaging to guide needles, wires and catheters using tiny access incisions. Like other minimally invasive techniques, risk, pain and recovery times are reduced as compared with more invasive approaches such as open surgery. These benefits, alongside the emergence of increasingly novel therapeutic technologies, are driving worldwide expansion. The core skills of IR include the Seldinger technique and the use of imaging and touch to effectively guide needles, wires and catheters in a wide range of procedures. Safe practice requires the operator to respond correctly to both visual and tactile cues in vascular angioplasty, stenting and stent-grafting, as well as control of bleeding, biopsy, abscess drainage and catheterization of the urinary and biliary tracts for drainage and stenting. The operator's deliberations may then initiate and inform a range of motor actions, including very fine translational and rotational motions, particularly in challenging anatomy. As the spectrum of available techniques increases, so the limited number and availability of suitably trained practitioners becomes a factor in their restricted availability to patients. Awareness of the need to expand IR training facilities is thus highly topical. |
International Conferences (peer reviewed articles)
| [19] |
 Zainab Ali Abbood and Franck P. Vidal.
Basic, dual, adaptive, and directed mutation operators in the Fly
algorithm.
In É. Lutton, P. Legrand, P. Parrend, N. Monmarché, and
M. Schoenauer, editors, Biennial International Conference on Artificial
Evolution (EA-2017), volume ???? of Lecture Notes in Computer Science,
pages ???--???, Paris, France, ??? 2018. Springer, Heidelberg.
Zainab Ali Abbood and Franck P. Vidal.
Basic, dual, adaptive, and directed mutation operators in the Fly
algorithm.
In É. Lutton, P. Legrand, P. Parrend, N. Monmarché, and
M. Schoenauer, editors, Biennial International Conference on Artificial
Evolution (EA-2017), volume ???? of Lecture Notes in Computer Science,
pages ???--???, Paris, France, ??? 2018. Springer, Heidelberg.



Our work is based on a Cooperative Co-evolution Algorithm -- the Fly algorithm -- in which individuals correspond to 3-D points. The Fly algorithm uses two levels of fitness function: i) a local fitness computed to evaluate a given individual (usually during the selection process) and ii) a global fitness to assess the performance of the population as a whole. This global fitness is the metrics that is minimised (or maximised depending on the problem) by the optimiser. Here the solution of the optimisation problem corresponds to a set of individuals instead of a single individual (the best individual) as in classical evolutionary algorithms. The Fly algorithm heavily relies on mutation operators and a new blood operator to insure diversity in the population. To lead to accurate results, a large mutation variance is often initially used to avoid local minima (or maxima). It is then progressively reduced to refine the results. Another approach is the use of adaptive operators. However, very little research on adaptive operators in Fly algorithm has been conducted. We address this deficiency and propose 4 different fully adaptive mutation operators in the Fly algorithm: positrons, and the final solution of the algorithm approximates the radioactivity concentration. The view and analysis four mutation operators, which are Basic Mutation, Adaptive Mutation Variance, Dual Mutation, and Directed Mutation. Due to the complex nature of the search space, (kN-dimensions, with k the number of genes per individuals and N the number of individuals in the population), we favour operators with a low maintenance cost in terms of computations. Their impact on the algorithm efficiency is analysed and validated on positron emission tomography (PET) reconstruction. Keywords: evolutionary algorithms, Parisian approach, reconstruction algorithms, positron emission tomography, mutation operator |
| [18] |
 Zainab Ali Abbood and Franck P. Vidal.
Basic, dual, adaptive, and directed mutation operators in the Fly
algorithm.
In É. Lutton, P. Legrand, P. Parrend, N. Monmarché, and
M. Schoenauer, editors, Biennial International Conference on Artificial
Evolution (EA-2017), pages 106--119, Paris, France, October 2017.
Zainab Ali Abbood and Franck P. Vidal.
Basic, dual, adaptive, and directed mutation operators in the Fly
algorithm.
In É. Lutton, P. Legrand, P. Parrend, N. Monmarché, and
M. Schoenauer, editors, Biennial International Conference on Artificial
Evolution (EA-2017), pages 106--119, Paris, France, October 2017.


Our work is based on a Cooperative Co-evolution Algorithm -- the Fly algorithm -- in which individuals correspond to 3-D points. The Fly algorithm uses two levels of fitness function: i) a local fitness computed to evaluate a given individual (usually during the selection process) and ii) a global fitness to assess the performance of the population as a whole. This global fitness is the metrics that is minimised (or maximised depending on the problem) by the optimiser. Here the solution of the optimisation problem corresponds to a set of individuals instead of a single individual (the best individual) as in classical evolutionary algorithms. The Fly algorithm heavily relies on mutation operators and a new blood operator to insure diversity in the population. To lead to accurate results, a large mutation variance is often initially used to avoid local minima (or maxima). It is then progressively reduced to refine the results. Another approach is the use of adaptive operators. However, very little research on adaptive operators in Fly algorithm has been conducted. We address this deficiency and propose 4 different fully adaptive mutation operators in the Fly algorithm: positrons, and the final solution of the algorithm approximates the radioactivity concentration. The view and analysis four mutation operators, which are Basic Mutation, Adaptive Mutation Variance, Dual Mutation, and Directed Mutation. Due to the complex nature of the search space, (kN-dimensions, with k the number of genes per individuals and N the number of individuals in the population), we favour operators with a low maintenance cost in terms of computations. Their impact on the algorithm efficiency is analysed and validated on positron emission tomography (PET) reconstruction. Keywords: evolutionary algorithms, Parisian approach, reconstruction algorithms, positron emission tomography, mutation operator |
| [17] |
 Zainab Ali Abbood and Franck P. Vidal.
Fly4arts: Evolutionary digital art with the fly algorithm.
In É. Lutton, P. Legrand, P. Parrend, N. Monmarché, and
M. Schoenauer, editors, Biennial International Conference on Artificial
Evolution (EA-2017), page 313, Paris, France, October 2017.
Zainab Ali Abbood and Franck P. Vidal.
Fly4arts: Evolutionary digital art with the fly algorithm.
In É. Lutton, P. Legrand, P. Parrend, N. Monmarché, and
M. Schoenauer, editors, Biennial International Conference on Artificial
Evolution (EA-2017), page 313, Paris, France, October 2017.


The aim of this study is to generate artistic images, such as digital mosaics, as an optimisation problem without the introduction of any a priori knowledge or constraint other than an input image. The usual practice to produce digital mosaic images heavily relies on Centroidal Voronoi diagrams. We demonstrate here that it can be modelled as an optimisation problem solved using a cooperative co-evolution strategy based on the Parisian evolution approach, the Fly algorithm. An individual is called a fly. Its aim of the algorithm is to optimise the position of innitely small 3-D points (the flies). The Fly algorithm has been initially used in real-time stereo vision for robotics. It has also demonstrated promising results in image reconstruction for tomography. In this new application, a much more complex representation has been studied. A fly is a tile. It has its own position, size, colour, and rotation angle. Our method takes advantage of graphics processing units (GPUs) to generate the images using the modern OpenGL Shading Language (GLSL) and Open Computing Language (OpenCL) to compute the difference between the input image and simulated image. Different types of tiles are implemented, some with transparency, to generate different visual effects, such as digital mosaic and spray paint. An online study with 41 participants has been conducted to compare some of our results with those generated using an open-source software for image manipulation. It demonstrates that our method leads to more visually appealing images. Keywords: Digital mosaic, Evolutionary art, Fly algorithm, Parisian evolution, cooperative co-evolution |
| [16] |
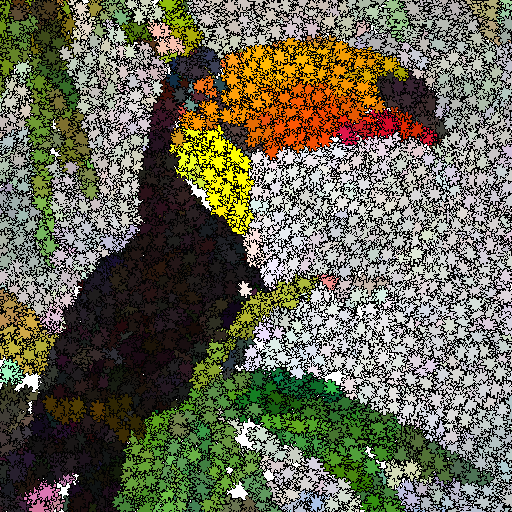 Z. Ali Abbood, O. Amlal, and F. P. Vidal.
Evolutionary Art Using the Fly Algorithm.
In Applications of Evolutionary Computation, volume 10199 of
Lecture Notes in Computer Science, pages 455-470, Amsterdam, The Netherlands,
April 2017. Springer, Heidelberg.
Z. Ali Abbood, O. Amlal, and F. P. Vidal.
Evolutionary Art Using the Fly Algorithm.
In Applications of Evolutionary Computation, volume 10199 of
Lecture Notes in Computer Science, pages 455-470, Amsterdam, The Netherlands,
April 2017. Springer, Heidelberg.



This study is about Evolutionary art such as digital mosaics. The most common techniques to generate a digital mosaic effect heavily rely on Centroidal Voronoi diagrams. Our method generates artistic images as an optimisation problem without the introduction of any a priori knowledge or constraint other than the input image. We adapt a cooperative co-evolution strategy based on the Parisian evolution approach, the Fly algorithm, to produce artistic visual effects from an input image (e.g. a photograph). The primary usage of the Fly algorithm is in computer vision, especially stereo-vision in robotics. It has also been used in image reconstruction for tomography. Until now the individuals correspond to simplistic primitives: Infinitely small 3-D points. In this paper, the individuals have a much more complex representation and represent tiles in a mosaic. They have their own position, size, colour, and rotation angle. We take advantage of graphics processing units (GPUs) to generate the images using the modern OpenGL Shading Language. Different types of tiles are implemented, some with transparency, to generate different visual effects, such as digital mosaic and spray paint. A user study has been conducted to evaluate some of our results. We also compare results with those obtained with GIMP, an open-source software for image manipulation. Keywords: Digital mosaic; Evolutionary art; Fly algorithm; Parisian evolution; Cooperative co-evolution |
| [15] |



Visual computing represents one of the most challenging and inspiring arenas in computer science. Today, fifty percent of content on the internet is in the form of visual data and information, and more than fifty percent of the neurons in the human brain are used in visual perception and reasoning. RIVIC is the collaborative amalgamation of research programmes between the computer science departments in Aberystwyth, Bangor, Cardiff and Swansea universities. Its aim is to promote research in visual computing, e.g. visualisation, computer vision and image/video processing, computer graphics and virtual environments. |
| [14] |


This paper shows new resutls of our artificial evolution algorithm for Positron Emission Tomography (PET) reconstruction. This imaging technique produces datasets corresponding to the concentration of positron emitters within the patient. Fully three-dimensional (3D) tomographic reconstruction requires high computing power and leads to many challenges. Our aim is to produce high quality datasets in a time that is clinically acceptable. Our method is based on a co-evolution strategy called the “Fly algorithm”. Each fly represents a point in space and mimics a positron emitter. Each fly position is progressively optimised using evolutionary computing to closely match the data measured by the imaging system. The performance of each fly is assessed based on its positive or negative contribution to the performance of the whole population. The final population of flies approximates the radioactivity concentration. This approach has shown promising results on numerical phantom models. The size of objects and their relative concentrations can be calculated in two-dimensional (2D) space. In (3D), complex shapes can be reconstructed. In this paper, we demonstrate the ability of the algorithm to fidely reconstruct more anatomically realistic volumes. Keywords: Evolutionary computation, inverse problems, adaptive algorithm, Nuclear medicine, Positron emission tomography, Reconstruction algorithms |
| [13] |


This paper is an overview of a method recently published in a biomedical journal (IEEE Transactions on Biomedical Engineering, http://tbme.embs.org). The method is based on an optimisation technique called “evolutionary strategy” and it has been designed to estimate the parameters of a complex 15-D respiration model. This model is adaptable to account for patient's specificities. The aim of the optimisation algorithm is to finely tune the model so that it accurately fits real patient datasets. The final results can then be embedded, for example, in high fidelity simulations of the human physiology. Our algorithm is fully automatic and adaptive. A compound fitness function has been designed to take into account for various quantities that have to be minimised (here topological errors of the liver and the diaphragm geometries). The performance our implementation is compared with two traditional methods (downhill simplex and conjugate gradient descent), a random search and a basic real-valued genetic algorithm. It shows that our evolutionary scheme provides results that are significantly more stable and accurate than the other tested methods. The approach is relatively generic and can be easily adapted to other complex parametrisation problems when ground truth data is available. Keywords: Evolutionary computation, inverse problems, medical simulation, adaptive algorithm |
| [12] |
 P.-F. Villard, F. P. Vidal, F. Bello, and N. W. John.
A method to compute respiration parameters for patient-based
simulators.
In Proceeding of Medicine Meets Virtual Reality 19 - NextMed
(MMVR19), volume 173 of Studies in Health Technology and Informatics,
pages 529-533, Newport Beach, California, February 2012. IOS Press.
Winner of the best poster award.
P.-F. Villard, F. P. Vidal, F. Bello, and N. W. John.
A method to compute respiration parameters for patient-based
simulators.
In Proceeding of Medicine Meets Virtual Reality 19 - NextMed
(MMVR19), volume 173 of Studies in Health Technology and Informatics,
pages 529-533, Newport Beach, California, February 2012. IOS Press.
Winner of the best poster award.



We propose a method to automatically tune a patient-based virtual environment training simulator for abdominal needle insertion. The key attributes to be customized in our framework are the elasticity of soft-tissues and the respiratory model parameters. The estimation is based on two 3D Computed Tomography (CT) scans of the same patient at two different time steps. Results are presented on five patients and show that our new method leads to better results than our previous studies with manually tuned parameters. |
| [11] |
 F. P. Vidal, É. Lutton, J. Louchet, and J.-M. Rocchisani.
Threshold selection, mitosis and dual mutation in cooperative
coevolution: application to medical 3D tomography.
In International Conference on Parallel Problem Solving From
Nature (PPSN'10), volume 6238 of Lecture Notes in Computer Science,
pages 414-423, Krakow, Poland, September 2010. Springer, Heidelberg.
F. P. Vidal, É. Lutton, J. Louchet, and J.-M. Rocchisani.
Threshold selection, mitosis and dual mutation in cooperative
coevolution: application to medical 3D tomography.
In International Conference on Parallel Problem Solving From
Nature (PPSN'10), volume 6238 of Lecture Notes in Computer Science,
pages 414-423, Krakow, Poland, September 2010. Springer, Heidelberg.



We present and analyse the behaviour of specialised operators designed for cooperative coevolution strategy in the framework of 3D tomographic PET reconstruction. The basis is a simple cooperative co-evolution scheme (the “fly algorithm”), which embeds the searched solution in the whole population, letting each individual be only a part of the solution. An individual, or fly, is a 3D point that emits positrons. Using a cooperative co-evolution scheme to optimize the position of positrons, the population of flies evolves so that the data estimated from flies matches measured data. The final population approximates the radioactivity concentration. In this paper, three operators are proposed, threshold selection, mitosis and dual mutation, and their impact on the algorithm efficiency is experimentally analysed on a controlled test-case. Their extension to other cooperative co-evolution schemes is discussed. |
| [10] |
 F. P. Vidal, M. Garnier, N. Freud, J. M. Létang, and N. W. John.
Accelerated deterministic simulation of x-ray attenuation using
graphics hardware.
In Eurographics 2010 - Poster, page Poster 5011, Norrköping,
Sweden, May 2010. Eurographics Association.
F. P. Vidal, M. Garnier, N. Freud, J. M. Létang, and N. W. John.
Accelerated deterministic simulation of x-ray attenuation using
graphics hardware.
In Eurographics 2010 - Poster, page Poster 5011, Norrköping,
Sweden, May 2010. Eurographics Association.


In this paper, we propose a deterministic simulation of X-ray transmission imaging on graphics hardware. Only the directly transmitted photons are simulated, using the Beer-Lambert law. Our previous attempt to simulate Xray attenuation from polygon meshes utilising the GPU showed significant increase of performance, with respect to a validated software implementation, without loss of accuracy. However, the simulations were restricted to monochromatic X-rays and finite point sources. We present here an extension to our method to perform physically more realistic simulations by taking into account polychromatic X-rays and focal spots causing blur. Keywords: Three-Dimensional Graphics and Realism; Raytracing; Physical Sciences and Engineering; Physics |
| [9] |
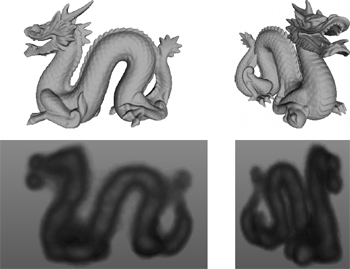 F. P. Vidal, J. Louchet, J.-M. Rocchisani, and É. Lutton.
New genetic operators in the Fly algorithm: application to medical
PET image reconstruction.
In Applications of Evolutionary Computation, volume 6024 of
Lecture Notes in Computer Science, pages 292-301, Istanbul, Turkey,
April 2010. Springer, Heidelberg.
Nominated for best paper award.
F. P. Vidal, J. Louchet, J.-M. Rocchisani, and É. Lutton.
New genetic operators in the Fly algorithm: application to medical
PET image reconstruction.
In Applications of Evolutionary Computation, volume 6024 of
Lecture Notes in Computer Science, pages 292-301, Istanbul, Turkey,
April 2010. Springer, Heidelberg.
Nominated for best paper award.



This paper presents an evolutionary approach for image reconstruction in positron emission tomography (PET). Our reconstruction method is based on a cooperative coevolution strategy (also called Parisian evolution): the “fly algorithm”. Each fly is a 3D point that mimics a positron emitter. The flies' position is progressively optimised using evolutionary computing to closely match the data measured by the imaging system. The performance of each fly is assessed using a “marginal evaluation” based on the positive or negative contribution of this fly to the performance of the population. Using this property, we propose a “thresholded-selection” method to replace the classical tournament method. A mitosis operator is also proposed. It is triggered to automatically increase the population size when the number of flies with negative fitness becomes too low. |
| [8] |
 F. P. Vidal, D. Lazaro-Ponthus, S. Legoupil, J. Louchet, É. Lutton, and
J.-M. Rocchisani.
Artificial evolution for 3D PET reconstruction.
In Proceedings of the 9th international conference on Artificial
Evolution (EA'09), volume 5975 of Lecture Notes in Computer Science,
pages 37-48, Strasbourg, France, October 2009. Springer, Heidelberg.
F. P. Vidal, D. Lazaro-Ponthus, S. Legoupil, J. Louchet, É. Lutton, and
J.-M. Rocchisani.
Artificial evolution for 3D PET reconstruction.
In Proceedings of the 9th international conference on Artificial
Evolution (EA'09), volume 5975 of Lecture Notes in Computer Science,
pages 37-48, Strasbourg, France, October 2009. Springer, Heidelberg.



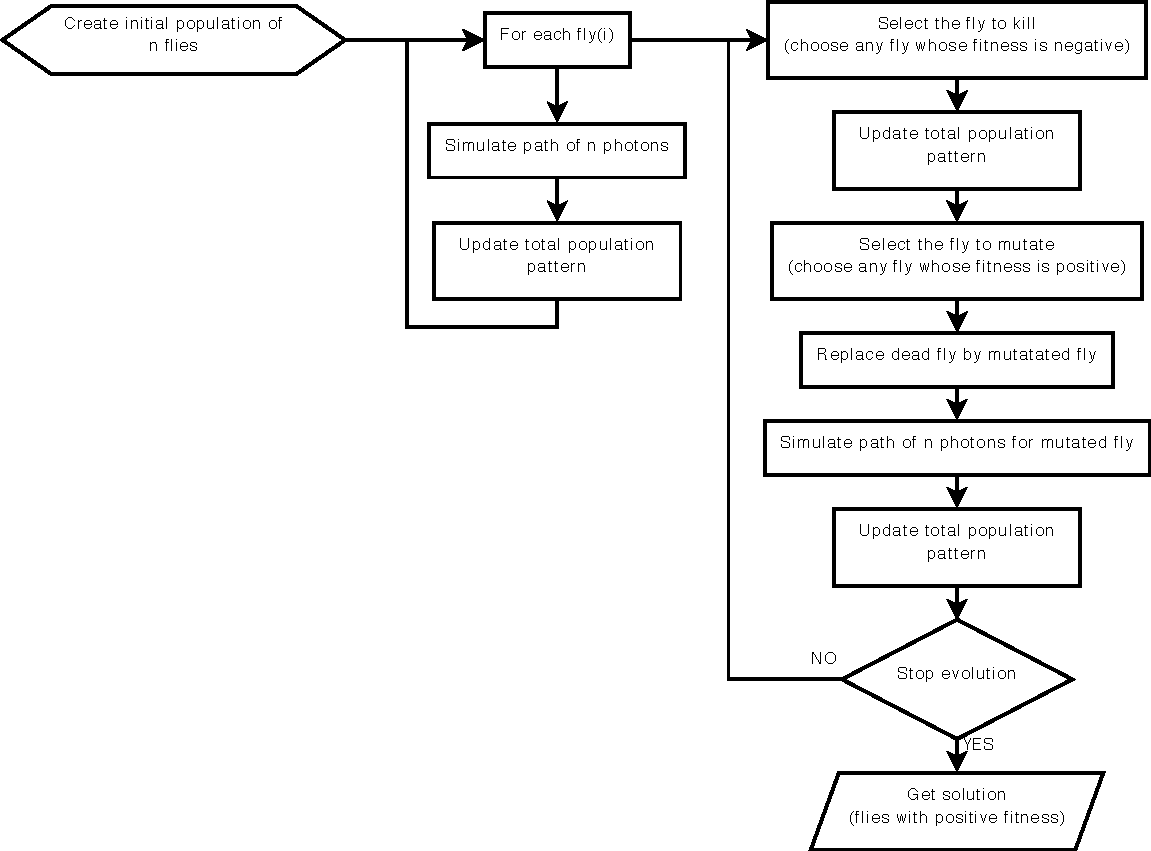
This paper presents a method to take advantage of artificial evolution in positron emission tomography reconstruction. This imaging technique produces datasets that correspond to the concentration of positron emitters through the patient. Fully 3D tomographic reconstruction requires high computing power and leads to many challenges. Our aim is to reduce the computing cost and produce datasets while retaining the required quality. Our method is based on a coevolution strategy (also called Parisian evolution) named “Fly algorithm”. Each fly represents a point of the space and acts as a positron emitter. The final population of flies corresponds to the reconstructed data. Using “marginal evaluation”, the fly's fitness is the positive or negative contribution of this fly to the performance of the population. This is also used to skip the relatively costly step of selection and simplify the evolutionary algorithm. |
| [7] |
 F. Bello, A. Bulpitt, D. A. Gould, R. Holbrey, C. Hunt, N. W. John, S. Johnson,
R. Phillips, A. Sinha, F. P. Vidal, P.-F. Villard, and H. Woolnough.
ImaGiNe-S: Imaging guided needle simulation.
In Eurographics 2009 - Medical Prize, pages 5-8, Munich,
Germany, March 2009. Eurographics Association.
Second prize and winner of 300.
F. Bello, A. Bulpitt, D. A. Gould, R. Holbrey, C. Hunt, N. W. John, S. Johnson,
R. Phillips, A. Sinha, F. P. Vidal, P.-F. Villard, and H. Woolnough.
ImaGiNe-S: Imaging guided needle simulation.
In Eurographics 2009 - Medical Prize, pages 5-8, Munich,
Germany, March 2009. Eurographics Association.
Second prize and winner of 300.



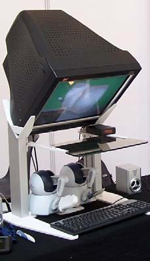
We present an integrated system for training visceral needle puncture procedures. Our aim is to provide a cost effective and validated training tool that uses actual patient data to enable interventional radiology trainees to learn how to carry out image-guided needle puncture. The input data required is a computed tomography scan of the patient that is used to create the patient specific models. Force measurements have been made on real tissue and the resulting data is incorporated into the simulator. Respiration and soft tissue deformations are also carried out to further improve the fidelity of the simulator. Keywords: Physically based modelling, Virtual reality |
| [6] |
 F. P. Vidal, P.-F. Villard, R. Holbrey, N. W. John, F. Bello, A. Bulpitt, and
D. A. Gould.
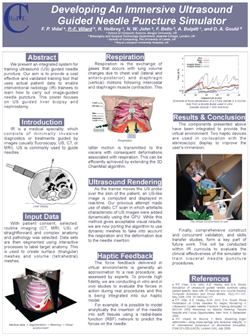
Developing an immersive ultrasound guided needle puncture simulator.
In Proceeding of Medicine Meets Virtual Reality 17 (MMVR17),
volume 142 of Studies in Health Technology and Informatics, pages
398-400, Long Beach, California, January 2009. IOS Press.
F. P. Vidal, P.-F. Villard, R. Holbrey, N. W. John, F. Bello, A. Bulpitt, and
D. A. Gould.
Developing an immersive ultrasound guided needle puncture simulator.
In Proceeding of Medicine Meets Virtual Reality 17 (MMVR17),
volume 142 of Studies in Health Technology and Informatics, pages
398-400, Long Beach, California, January 2009. IOS Press.



We present an integrated system for training ultrasound guided needle puncture. Our aim is to provide a cost effective and validated training tool that uses actual patient data to enable interventional radiology trainees to learn how to carry out image-guided needle puncture. The input data required is a computed tomography scan of the patient that is used to create the patient specific models. Force measurements have been made on real tissue and the resulting data is incorporated into the simulator. Respiration and soft tissue deformations are also carried out to further improve the fidelity of the simulator. Keywords: image guided needle puncture training, interventional radiology training, needle puncture |
| [5] |
 L. ap Cynydd, N. W. John, F. P. Vidal, D. A. Gould, E. Joekes, and
P. Littler.
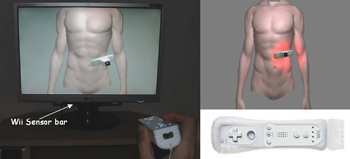
Cost effective ultrasound imaging training mentor for use in
developing countries.
In Proceeding of Medicine Meets Virtual Reality 17 (MMVR17),
volume 142 of Studies in Health Technology and Informatics, pages
49-54, Long Beach, California, January 2009. IOS Press.
L. ap Cynydd, N. W. John, F. P. Vidal, D. A. Gould, E. Joekes, and
P. Littler.
Cost effective ultrasound imaging training mentor for use in
developing countries.
In Proceeding of Medicine Meets Virtual Reality 17 (MMVR17),
volume 142 of Studies in Health Technology and Informatics, pages
49-54, Long Beach, California, January 2009. IOS Press.



This paper reports on a low cost system for training ultrasound imaging techniques. The need for such training is particularly acute in developing countries where typically ultrasound scanners remain idle due to the lack of experienced sonographers. The system described below is aimed at a PC platform but uses interface components from the Nintendo Wii games console. The training software is being designed to support a variety of patient case studies, and also supports remote tutoring over the internet. Keywords: Ultrasound Training, medical virtual environment, hci |
| [4] |
 N. W. John, V. Luboz, F. Bello, C. Hughes, F. P. Vidal, I. S. Lim, T. V. How,
J. Zhai, S. Johnson, N. Chalmers, K. Brodlie, A. Bulpit, Y. Song, D. O.
Kessel, R. Phillips, J. W. Ward, S. Pisharody, Y. Zhang, C. M. Crawshaw, and
D. A. Gould.
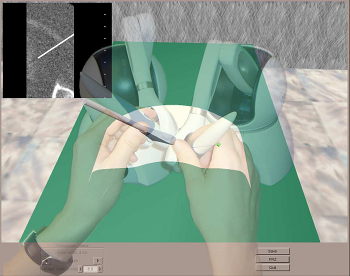
Physics-based virtual environment for training core skills in
vascular interventional radiological procedures.
In Proceeding of Medicine Meets Virtual Reality 16 (MMVR16),
volume 132 of Studies in Health Technology and Informatics, pages
195-197, Long Beach, California, January 2008. IOS Press.
N. W. John, V. Luboz, F. Bello, C. Hughes, F. P. Vidal, I. S. Lim, T. V. How,
J. Zhai, S. Johnson, N. Chalmers, K. Brodlie, A. Bulpit, Y. Song, D. O.
Kessel, R. Phillips, J. W. Ward, S. Pisharody, Y. Zhang, C. M. Crawshaw, and
D. A. Gould.
Physics-based virtual environment for training core skills in
vascular interventional radiological procedures.
In Proceeding of Medicine Meets Virtual Reality 16 (MMVR16),
volume 132 of Studies in Health Technology and Informatics, pages
195-197, Long Beach, California, January 2008. IOS Press.



Recent years have seen a significant increase in the use of Interventional Radiology (IR) as an alternative to open surgery. A large number of IR procedures commences with needle puncture of a vessel to insert guidewires and catheters: these clinical skills are acquired by all radiologists during training on patients, associated with some discomfort and occasionally, complications. While some visual skills can be acquired using models such as the ones used in surgery, these have limitations for IR which relies heavily on a sense of touch. Both patients and trainees would benefit from a virtual environment (VE) conveying touch sensation to realistically mimic procedures. The authors are developing a high fidelity VE providing a validated alternative to the traditional apprenticeship model used for teaching the core skills. The current version of the CRaIVE simulator combines home made software, haptic devices and commercial equipments. Keywords: Virtual environment, patient specific model, interventional radiology |
| [3] |
 F. P. Vidal, N. W. John, and R.M. Guillemot.
Interactive physically-based X-ray simulation: CPU or GPU?
In Proceeding of Medicine Meets Virtual Reality 15 (MMVR15),
volume 125 of Studies in Health Technology and Informatics, pages
479-481, Long Beach, California, February 2007. IOS Press.
F. P. Vidal, N. W. John, and R.M. Guillemot.
Interactive physically-based X-ray simulation: CPU or GPU?
In Proceeding of Medicine Meets Virtual Reality 15 (MMVR15),
volume 125 of Studies in Health Technology and Informatics, pages
479-481, Long Beach, California, February 2007. IOS Press.


Interventional Radiology (IR) procedures are minimally invasive, targeted treatments performed using imaging for guidance. Needle puncture using ultrasound, x-ray, or computed tomography (CT) images is a core task in the radiology curriculum, and we are currently devel- oping a training simulator for this. One requirement is to include support for physically-based simulation of x-ray images from CT data sets. In this paper, we demonstrate how to exploit the capability of today's graphics cards to efficiently achieve this on the Graphics Processing Unit (GPU) and compare performance with an efficient software only implementation using the Central Processing Unit (CPU). Keywords: X-ray simulation, GPU-based volume rendering, Interventional radiology |
| [2] |
 F. P. Vidal, N. Chalmers, D. A. Gould, A. E. Healey, and N. W. John.
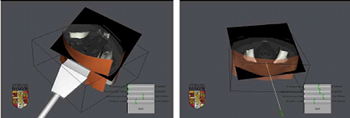
Developing a needle guidance virtual environment with patient
specific data and force feedback.
In Proceeding of the 19th International Congress of Computer
Assisted Radiology and Surgery (CARS'05), volume 1281 of International
Congress Series, pages 418-423, Berlin, Germany, June 2005. Elsevier.
F. P. Vidal, N. Chalmers, D. A. Gould, A. E. Healey, and N. W. John.
Developing a needle guidance virtual environment with patient
specific data and force feedback.
In Proceeding of the 19th International Congress of Computer
Assisted Radiology and Surgery (CARS'05), volume 1281 of International
Congress Series, pages 418-423, Berlin, Germany, June 2005. Elsevier.



We present a simulator for guided needle puncture procedures. Our aim is to provide an effective training tool for students in interventional radiology (IR) using actual patient data and force feedback within an immersive virtual environment (VE). Training of the visual and motor skills required in IR is an apprenticeship which still consists of close supervision using the model: (i) see one, (ii) do one, and (iii) teach one. Training in patients not only has discomfort associated with it, but provides limited access to training scenarios, and makes it difficult to train in a time efficient manner. Currently, the majority of commercial products implementing a medical VE still focus on laparoscopy where eye-hand coordination and sensation are key issues. IR procedures, however, are far more reliant on the sense of touch. Needle guidance using ultrasound or computed tomography (CT) images is also widely used. Both of these are areas that have not been fully addressed by other medical VEs. This paper provides details of how we are developing an effective needle guidance simulator. The project is a multi-disciplinary collaboration involving practising interventional radiologists and computer scientists. Keywords: Interventional radiology; Virtual environments; Needle puncture; Haptics |
| [1] |
 F. P. Vidal, F. Bello, K. Brodlie, R. Phillips N. W. John, D. Gould, and
N. Avis.
Principles and applications of medical virtual environments.
In Christophe Schlick and Werner Purgathofer, editors,
State-of-the-art Proceedings of Eurographics 2004, pages 1-35, Grenoble,
France, August 2004. Eurographics Association.
F. P. Vidal, F. Bello, K. Brodlie, R. Phillips N. W. John, D. Gould, and
N. Avis.
Principles and applications of medical virtual environments.
In Christophe Schlick and Werner Purgathofer, editors,
State-of-the-art Proceedings of Eurographics 2004, pages 1-35, Grenoble,
France, August 2004. Eurographics Association.



The medical domain offers many excellent opportunities for the application of computer graphics, visualization, and virtual environments, offering the potential to help improve healthcare and bring benefits to patients. This report provides a comprehensive overview of the state-of-the-art in this exciting field. It has been written from the perspective of both computer scientists and practicing clinicians and documents past and current successes together with the challenges that lie ahead. The report begins with a description of the commonly used imaging modalities and then details the software algorithms and hardware that allows visualization of and interaction with this data. Example applications from research projects and commercially available products are listed, including educational tools; diagnostic aids; virtual endoscopy; planning aids; guidance aids; skills training; computer augmented reality; and robotics. The final section of the report summarises the current issues and looks ahead to future developments. Keywords: Augmented and virtual realities, Computer Graphics, Health, Physically based modelling, Medical Sciences, Simulation, Virtual device interfaces |
International Conferences (invited oral presentation)
| [1] |
F. P. Vidal.
Developing a virtual environment for training in visceral
interventional radiology.
In Workshop on Open Source Haptics & Applications, EuroHaptics
2008 (EH 2008), Madrid, Spain, February 2008.
Invited talk.


|
International Conferences (abstracts)
| [1] |
 F. P. Vidal, M. Folkerts, N. Freud, and S. Jiang.
GPU accelerated DRR computation with scatter.
Medical Physics, 38(6):3455-3456, July 2011.
F. P. Vidal, M. Folkerts, N. Freud, and S. Jiang.
GPU accelerated DRR computation with scatter.
Medical Physics, 38(6):3455-3456, July 2011.


Purpose: We propose a fast software library implemented on
graphics processing unit (GPU) to compute digitally reconstructed
radiographs (DRRs). It takes into account first order Compton
scattering. |
| [2] |
 F. P. Vidal, J. Louchet, J.-M. Rocchisani, and É. Lutton.
Flies for PET: An artificial evolution strategy for image
reconstruction in nuclear medicine.
Medical Physics, 37(6):3139, July 2010.
F. P. Vidal, J. Louchet, J.-M. Rocchisani, and É. Lutton.
Flies for PET: An artificial evolution strategy for image
reconstruction in nuclear medicine.
Medical Physics, 37(6):3139, July 2010.



Purpose: We propose an evolutionary approach for image
reconstruction in nuclear medicine. Our method is based on
a cooperative coevolution strategy (also called Parisian evolution):
the “fly algorithm”. |
| [3] |
 F. P. Vidal, P. F. Villard, M. Garnier, N. Freud, J. M. Létang, N. W. John,
and F. Bello.
Joint simulation of transmission x-ray imaging on GPU and patient's
respiration on CPU.
Medical Physics, 37(6):3129, July 2010.
F. P. Vidal, P. F. Villard, M. Garnier, N. Freud, J. M. Létang, N. W. John,
and F. Bello.
Joint simulation of transmission x-ray imaging on GPU and patient's
respiration on CPU.
Medical Physics, 37(6):3129, July 2010.



Purpose: We previously proposed to compute the X‐ray attenuation
from polygons directly on the GPU, using OpenGL, to significantly
increase performance without loss of accuracy. The method has been
deployed into a training simulator for percutaneous transhepatic
cholangiography. The simulations were however restricted to
monochromatic X‐rays using a point source. They now take into account
both the geometrical blur and polychromatic X‐rays. |
| [4] |
A. Sinha, K. Flood, D. Kessel, S. Johnson, C. Hunt, H. Woolnough, F. P. Vidal,
P.-F. Villard, R. Holbray, C. M. Crawshaw, A. Bulpitt, N. W. John,
F. Bello, R. Phillips, and D. A. Gould.
The role of simulation in medical training and assessment.
In Radiological Society of North America 2009 (RSNA 2009),
Chicago, Illinois, November 2009.

|
| [5] |
 P.-F. Villard, F. P. Vidal, C. Hunt, F. Bello, N. W. John, S. Johnson, and
D. A. Gould.
Percutaneous transhepatic cholangiography training simulator with
real-time breathing motion.
In Proceeding of the 23rd International Congress of Computer
Assisted Radiology and Surgery, volume 4 (Suppl 1) of International
Journal of Computer Assisted Radiology and Surgery, pages S66-S67, Berlin,
Germany, June 2009. Springer.
P.-F. Villard, F. P. Vidal, C. Hunt, F. Bello, N. W. John, S. Johnson, and
D. A. Gould.
Percutaneous transhepatic cholangiography training simulator with
real-time breathing motion.
In Proceeding of the 23rd International Congress of Computer
Assisted Radiology and Surgery, volume 4 (Suppl 1) of International
Journal of Computer Assisted Radiology and Surgery, pages S66-S67, Berlin,
Germany, June 2009. Springer.


Keywords: Interventional radiology, Virtual environments, Respiration simulation, X-ray simulation, Needle puncture, Haptics, Task analysis |
International Conferences (other presentations)
| [1] |
 Zainab Ali Abbood, Jean-Marie Rocchisani, and Franck P. Vidal.
Visualisation of PET data in the Fly Algorithm.
In Katja Bühler, Lars Linsen, and Nigel W. John, editors,
Eurographics Workshop on Visual Computing for Biology and Medicine, pages
211--212. The Eurographics Association, 2015.
Zainab Ali Abbood, Jean-Marie Rocchisani, and Franck P. Vidal.
Visualisation of PET data in the Fly Algorithm.
In Katja Bühler, Lars Linsen, and Nigel W. John, editors,
Eurographics Workshop on Visual Computing for Biology and Medicine, pages
211--212. The Eurographics Association, 2015.

 | |

We use the Fly algorithm, an artificial evolution strategy, to reconstruct positron emission tomography (PET) images. The algorithm iteratively optimises the position of 3D points. It eventually produces a point cloud, which needs to be voxelised to produce volume data that can be used with conventional medical image software. However, resulting voxel data is noisy. In our test case with 6,400 points the normalised cross-correlation (NCC) between the reference and the reconstruction is 85.53%; with 25,600 points it is 93.60%. This paper introduces a more robust 3D voxelisation method based on implicit modelling using metaballs to overcome this limitation. With metaballs, the NCC with 6,400 points increases up to 92.21%; and up to 96.26% with 25,600 points. |
| [2] |
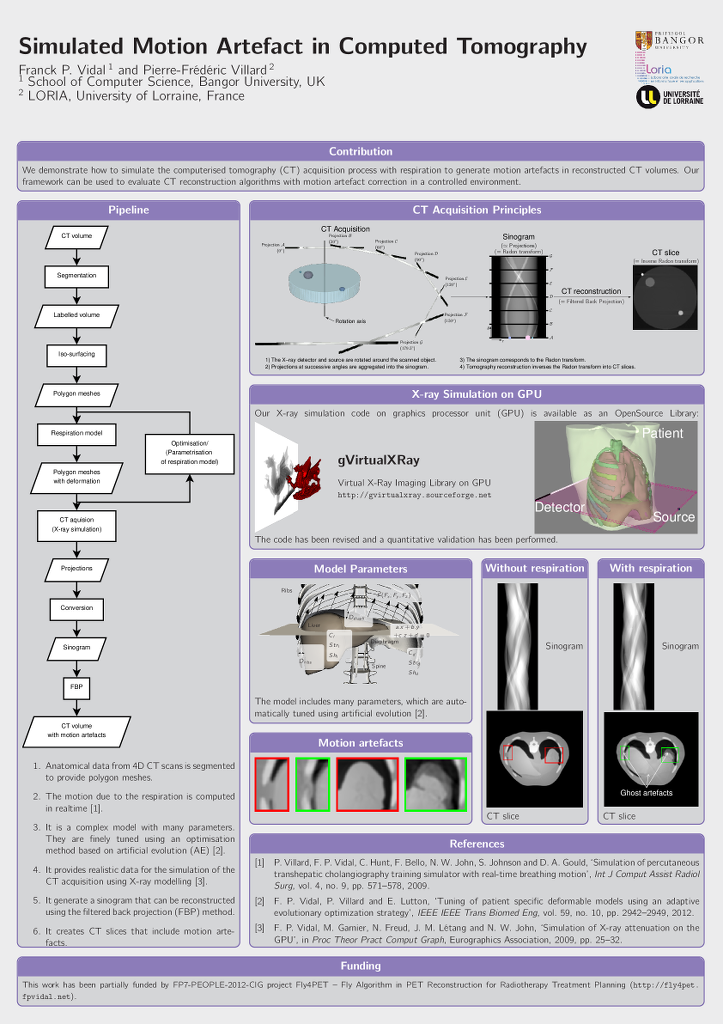 Franck P. Vidal and Pierre-Frédéric Villard.
Simulated Motion Artefact in Computed Tomography.
In Katja Bühler, Lars Linsen, and Nigel W. John, editors,
Eurographics Workshop on Visual Computing for Biology and Medicine, pages
213--214. The Eurographics Association, 2015.
Franck P. Vidal and Pierre-Frédéric Villard.
Simulated Motion Artefact in Computed Tomography.
In Katja Bühler, Lars Linsen, and Nigel W. John, editors,
Eurographics Workshop on Visual Computing for Biology and Medicine, pages
213--214. The Eurographics Association, 2015.

 | |

We propose a simulation framework to simulate the computed tomography acquisition process. It includes five components: anatomic data, respiration modelling, automatic parametrisation, X-ray simulation, and tomography reconstruction. It is used to generate motion artefacts in reconstructed CT volumes. Our framework can be used to evaluate CT reconstruction algorithm with motion artefact correction in a controlled environment. |
| [3] |
 F. Bello, T. R. Coles, D. A. Gould, C. J. Hughes, N. W. John, F. P. Vidal, and
S. Watt.
The need to touch medical virtual environments?
In IEEE Virtual Reality 2010 (VR2010), Workshop on Medical
Virtual Environments, Waltham, Massachusetts, March 2010. IEEE Computer
Society.
Available online at http://www.hpv.cs.bangor.ac.uk/vr10-med/.
F. Bello, T. R. Coles, D. A. Gould, C. J. Hughes, N. W. John, F. P. Vidal, and
S. Watt.
The need to touch medical virtual environments?
In IEEE Virtual Reality 2010 (VR2010), Workshop on Medical
Virtual Environments, Waltham, Massachusetts, March 2010. IEEE Computer
Society.
Available online at http://www.hpv.cs.bangor.ac.uk/vr10-med/.


Haptics technologies are frequently used in virtual environments to allow participants to touch virtual objects. Medical applications are no exception and a wide variety of commercial and bespoke haptics hardware solutions have been employed to aid in the simulation of medical procedures. Intuitively the use of haptics will improve the training of the task. However, little evidence has been published to prove that this is indeed the case. In the paper we summarise the available evidence and use a case study from interventional radiology to discuss the question of how important is it to touch medical virtual environments? Keywords: Haptics, virtual environments, touch |
| [4] |
 F. P. Vidal, J. Louchet, É. Lutton, and J.-M. Rocchisani.
PET reconstruction using a cooperative coevolution strategy in
LOR space.
In IEEE Nuclear Science Symposium Conference Record, pages
3363-3366, Orlando, Florida, October 2009. IEEE.
F. P. Vidal, J. Louchet, É. Lutton, and J.-M. Rocchisani.
PET reconstruction using a cooperative coevolution strategy in
LOR space.
In IEEE Nuclear Science Symposium Conference Record, pages
3363-3366, Orlando, Florida, October 2009. IEEE.


This paper presents preliminary results of a novel method that takes advantage of artificial evolution for positron emission tomography (PET) reconstruction. Fully 3D tomographic reconstruction in PET requires high computing power and leads to many challenges. To date, the use of such methods is still restricted due to the heavy computing power needed. Evolutionary algorithms have proven to be efficient optimisation techniques in various domains. However the use of evolutionary computation in tomographic reconstruction has been largely overlooked. We propose a computer-based algorithm for fully 3D reconstruction in PET based on artificial evolution and evaluate its relevance. Keywords: Positron emission tomography, genetic algorithms, optimization methods |
| [5] |
 N. W. John, C. Hughes, S. Pop, F. P. Vidal, and O. Buckley.
Computational requirements of the virtual patient.
In Proceedings of the First International Conference on
Computational and Mathematical Biomedical Engineering (CMBE 2009), pages
140-143, Swansea, UK, June 2009.
N. W. John, C. Hughes, S. Pop, F. P. Vidal, and O. Buckley.
Computational requirements of the virtual patient.
In Proceedings of the First International Conference on
Computational and Mathematical Biomedical Engineering (CMBE 2009), pages
140-143, Swansea, UK, June 2009.

Medical visualization in a hospital can be used to aid training, diagnosis, and pre- and intra-operative planning. In such an application, a virtual representation of a patient is needed that is interactive, can be viewed in three dimensions (3D), and simulates physiological processes that change over time. This paper highlights some of the computational challenges of implementing a real time simulation of a virtual patient, when accuracy can be traded-off against speed. Illustrations are provided using projects from our research based on Grid-based visualization, through to use of the Graphics Processing Unit (GPU). Keywords: Medical visualization, virtual environment, Grid, GPU |
| [6] |
D. A. Gould, F. P. Vidal, C. Hughes, P. F. Villard, V. Luboz, N. W. John,
F. Bello, A. Bulpitt, V. Gough, and D. O. Kessel.
Interventional radiology core skills simulation: mid term status of
the CRaIVE projects.
In Cardiovascular and Interventional Radiological Society of
Europe 2008 (CIRCE 2008), Electronic Poster, page P 130, Copenhagen,
Denmark, September 2008.

|
| [7] |
 F. P. Vidal, A. E. Healey, N. W. John, and D. A. Gould.
Force penetration of chiba needles for haptic rendering in ultrasound
guided needle puncture training simulator.
In MICCAI 2008 - Workshop on Needle Steering: Recent Results
and Future Opportunities, New York, September 2008.
Available at http://lcsr.jhu.edu/NeedleSteering/Workshop/Vidal.html.
F. P. Vidal, A. E. Healey, N. W. John, and D. A. Gould.
Force penetration of chiba needles for haptic rendering in ultrasound
guided needle puncture training simulator.
In MICCAI 2008 - Workshop on Needle Steering: Recent Results
and Future Opportunities, New York, September 2008.
Available at http://lcsr.jhu.edu/NeedleSteering/Workshop/Vidal.html.



|
| [8] |
 G. Debouzy, F. Vidal, D. Deprez, S. Keswani, J. Warren, and P. Cosson.
Virtual radiographic environment.
In Eurographics 2003 Medical Prize, Granada, Spain, September
2003.
G. Debouzy, F. Vidal, D. Deprez, S. Keswani, J. Warren, and P. Cosson.
Virtual radiographic environment.
In Eurographics 2003 Medical Prize, Granada, Spain, September
2003.


|
National Conferences
| [1] |
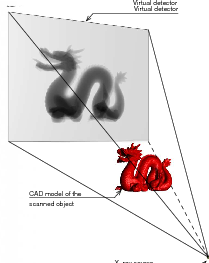 F. P. Vidal, M. Garnier, N. Freud, J. M. Létang, and N. W. John.
Simulation of X-ray attenuation on the GPU.
In Proceedings of Theory and Practice of Computer Graphics
2009, pages 25-32, Cardiff, UK, June 2009. Eurographics Association.
Winner of Ken Brodlie Prize for Best Paper.
F. P. Vidal, M. Garnier, N. Freud, J. M. Létang, and N. W. John.
Simulation of X-ray attenuation on the GPU.
In Proceedings of Theory and Practice of Computer Graphics
2009, pages 25-32, Cardiff, UK, June 2009. Eurographics Association.
Winner of Ken Brodlie Prize for Best Paper.



In this paper, we propose to take advantage of computer graphics hardware to achieve an accelerated simulation of X-ray transmission imaging, and we compare results with a fast and robust software-only implementation. The running times of the GPU and CPU implementations are compared in different test cases. The results show that the GPU implementation with full floating point precision is faster by a factor of about 60 to 65 than the CPU implementation, without any significant loss of accuracy. The increase in performance achieved with GPU calculations opens up new perspectives. Notably, it paves the way for physically-realistic simulation of X-ray imaging in interactive time. Keywords: Physically based modeling, Raytracing, Physics |
| [2] |
 A. Sinha, S. Johnson, C. Hunt, H. Woolnough, F. P. Vidal, and D. Gould.
Preliminary face and content validation of Imagine-S: the CIRSE
& BSIR experience.
In Proceedings of the UK Radiological Congress, page 2,
Manchester, UK, June 2009.
A. Sinha, S. Johnson, C. Hunt, H. Woolnough, F. P. Vidal, and D. Gould.
Preliminary face and content validation of Imagine-S: the CIRSE
& BSIR experience.
In Proceedings of the UK Radiological Congress, page 2,
Manchester, UK, June 2009.

KEY LEARNING OBJECTIVES:
To determine face and content validity of a physics-based virtual reality (VR)
training simulation of visceral interventional radiology needle puncture procedures. |
| [3] |
 P. F. Villard, P. Littler, V. Gough, F. P. Vidal, C. Hughes, N. W. John,
V. Luboz, F. Bello, Y. Song, R. Holbrey, A. Bulpitt, D. Mullan, N. Chalmers,
D. Kessel, and D. Gould.
Improving the modeling of medical imaging data for simulation.
In Proceedings of the UK Radiological Congress, page 61,
Birmingham, UK, June 2008.
P. F. Villard, P. Littler, V. Gough, F. P. Vidal, C. Hughes, N. W. John,
V. Luboz, F. Bello, Y. Song, R. Holbrey, A. Bulpitt, D. Mullan, N. Chalmers,
D. Kessel, and D. Gould.
Improving the modeling of medical imaging data for simulation.
In Proceedings of the UK Radiological Congress, page 61,
Birmingham, UK, June 2008.

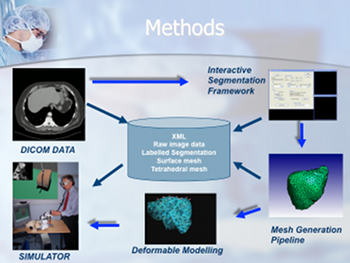
PURPOSE-MATERIALS:
To use patient imaging as the basis for developing virtual environments (VE). |
| [4] |
 P. Cosson, J. Yu Cheng, S. Keswani, G. Debouzy, D. Deprez, and F. Vidal.
Virtual radiographic environments become a reality.
In Proceedings of the UK Radiological Congress, Manchester, UK,
June 2004.
P. Cosson, J. Yu Cheng, S. Keswani, G. Debouzy, D. Deprez, and F. Vidal.
Virtual radiographic environments become a reality.
In Proceedings of the UK Radiological Congress, Manchester, UK,
June 2004.

|
| [5] |
P. Cosson, G. Debouzy, D. Deprez, F. Vidal, S. Keswani, and J. Warren.
Virtual radiographic environments: what use would they be?
In University of Teesside Annual Learning & Teaching
Conference, Middlesbrough, UK, January 2004.

|
| [6] |
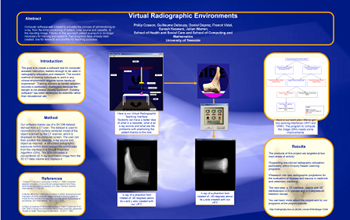 P. Cosson, G. Debouzy, D. Deprez, F. Vidal, S. Keswani, and J. Warren.
Virtual radiographic environments.
In University of Teesside Annual Learning & Teaching
Conference, Middlesbrough, UK, 2003.
P. Cosson, G. Debouzy, D. Deprez, F. Vidal, S. Keswani, and J. Warren.
Virtual radiographic environments.
In University of Teesside Annual Learning & Teaching
Conference, Middlesbrough, UK, 2003.

Computer software was created to simulate the process of administering an x-ray, from the initial positioning of patient, x-ray source and cassette, to the resulting image. Thanks to this approach patient exposure is no longer necessary for training and research. Two programs have already been created: one for research and another for teaching purposes |
Postgraduate Dissertations
| [1] |
 F. P. Vidal.
Simulation of image guided needle puncture: contribution to
real-time ultrasound and fluoroscopic rendering, and volume haptic
rendering.
PhD thesis, School of Computer Science, Bangor University, UK,
January 2008.
F. P. Vidal.
Simulation of image guided needle puncture: contribution to
real-time ultrasound and fluoroscopic rendering, and volume haptic
rendering.
PhD thesis, School of Computer Science, Bangor University, UK,
January 2008.


The potential for the use of computer graphics in medicine has been well established. An important emerging area is the provision of training tools for interventional radiology (IR) procedures. These are minimally invasive, targeted treatments performed using imaging for guidance. Training of the skills required in IR is an apprenticeship which still consists of close supervision using the model i) see one, ii) do one and iii) teach one. Simulations of guidewire and catheter insertion for IR are already commercially available. However, training of needle guidance using ultrasound (US), fluoroscopic or computed tomography (CT) images - the first step in approximately half of all IR procedures - has been largely overlooked and we have developed a simulator, called BIGNePSi, to provide training for this commonly performed procedure. This thesis is devoted to the development of novel techniques to provide an integrated visual-haptic system for the simulation of US guided needle puncture using patient specific data with 3D textures and volume haptics. The result is the realization of a cost effective training tool, using off-the-shelf components (visual displays, haptic devices and working stations), that delivers a high fidelity training experience. We demonstrate that the proxy-based haptic rendering method can be extended to use volumetric data so that the trainee can feel underlying structures, such as ribs and bones, whilst scanning the surface of the body with a virtual US transducer. A volume haptic model is also proposed that implements an effective model of needle puncture that can be modulated by using actual force measurements. A method of approximating US-like images from CT data sets is also described. We also demonstrate how to exploit today's graphics cards to achieve physically-based simulation of x-ray images using GPU programming and 3D texture hardware. We also demonstrate how to use GPU programming to modify, at interactive framerates, the content of 3D textures to include the needle shaft and also to artificially add a tissue lesion into the dataset of a specific patient. This enables the clinician to provide students with a wide variety of training scenarios. Validation of the simulator is critical to its eventual uptake in a training curriculum and a project such as this cannot be undertaken without close co-operation with the domain experts. Hence this project has been undertaken within a multi-disciplinary collaboration involving practising interventional radiologists and computer scientists of the Collaborators in Radiological Interventional Virtual Environments (CRaIVE) consortium. The cognitive task analysis (CTA) for freehand US guided biopsy performed by our psychologist partners has been extensively used to guide the design of the simulator. In addition, to ensure that the fidelity of the simulator is at an acceptable level, clinical validation of the system's content has been carried out at each development stage. In further, objective evaluations, questionnaires were developed to evaluate the features and the performances of the simulator. These were distributed to trainees and experts at different workshops. Many suggestions for improvements were collected and subsequently integrated into the simulator. |
| [2] |
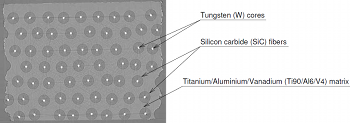 F. P. Vidal.
Modelling the response of x-ray detectors and removing artefacts in
3D tomography.
Master's thesis, École doctorale Électronique,
Électrotechnique, Automatique, INSA de Lyon, France, September 2003.
F. P. Vidal.
Modelling the response of x-ray detectors and removing artefacts in
3D tomography.
Master's thesis, École doctorale Électronique,
Électrotechnique, Automatique, INSA de Lyon, France, September 2003.


This work presents a method for modelling the response of X-ray detectors applied to remove artefacts in tomography. On some reconstructed volumes by tomography using synchrotron radiations at ESRF, dark line artefacts (under-estimation of linear attenuation coefficients) appear when high density material are aligned. The causes of these artefacts have been determined using experimental and simulation methods; then simulated artefacts were removed. Finally two different causes are highlighted in this study. One of them is the impulse response of the used detector, the Frelon camera of ID19 beamline. An iterative fixed point algorithm has been used successfully to remove simulated artefacts on tomographic slices. Keywords: tomography; X-ray; detectors response modelling; artefacts; simulation; deconvolution |
| [3] |
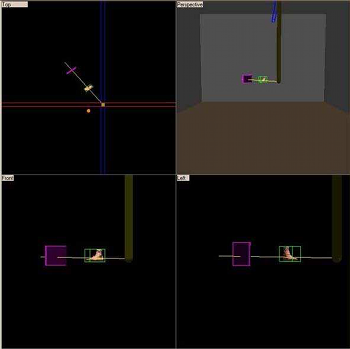 F. P. Vidal.
Constructing a GUI using 3D reconstruction for a radiographer's
training tool.
Master's thesis, School of Computing and Mathematics, University of
Teesside, UK, September 2002.
F. P. Vidal.
Constructing a GUI using 3D reconstruction for a radiographer's
training tool.
Master's thesis, School of Computing and Mathematics, University of
Teesside, UK, September 2002.


This project presents the GUI for a radiographer's training tool. It is one of the two parts of a project idea proposed by Senior Lecturer, Philip Cosson, from the School of Health and Social Care at the University of Teesside. His wish was to develop a program, which would allow students to train themselves to take X-Ray images without exposing patients to X-rays. It has been divided into two different projects, the GUI and the X-Ray rendering. There are two parts to the GUI system, a 3D reconstruction and the setting of the radiography parameters via the GUI. Volumetric data, obtained by MR or CT scanners, is stored, slice by slice, in a DICOM file, the medical imaging file standard. The Papyrus toolkit, developed at the University Hospital of Geneva, is used to read DICOMDIR files and DICOM files, which contain medical images. Before 3D reconstruction, information is extracted and a segmentation algorithm detects bones and skin on the different images of the dataset. From these data and using the marching cube algorithm, a 3D model is created. The GUI lets users select different datasets. Users can set the position and the orientation of the 3D reconstructed objects. They are able to choose the corresponding cassette and to move the X-Ray source. The GUI communicates, via an agreed protocol, all settings of the scene to the other part of the project, the X-Ray renderer. The GUI project is written in C++ using the WIN32 API and OpenGL. |
Parts of this file were generated by bibtex2html 1.97.

